Structural basis of lipoprotein recognition by the bacterial Lol
Por um escritor misterioso
Descrição

Lipoproteins in bacteria: structures and biosynthetic pathways - Nakayama - 2012 - The FEBS Journal - Wiley Online Library
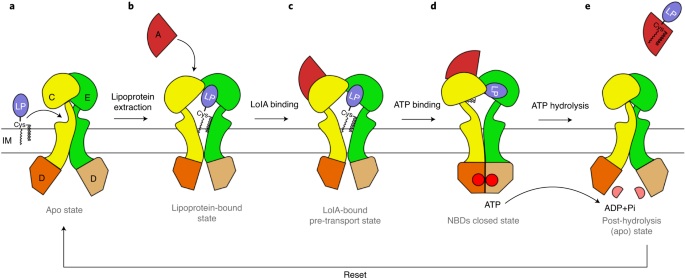
Structural basis for bacterial lipoprotein relocation by the transporter LolCDE
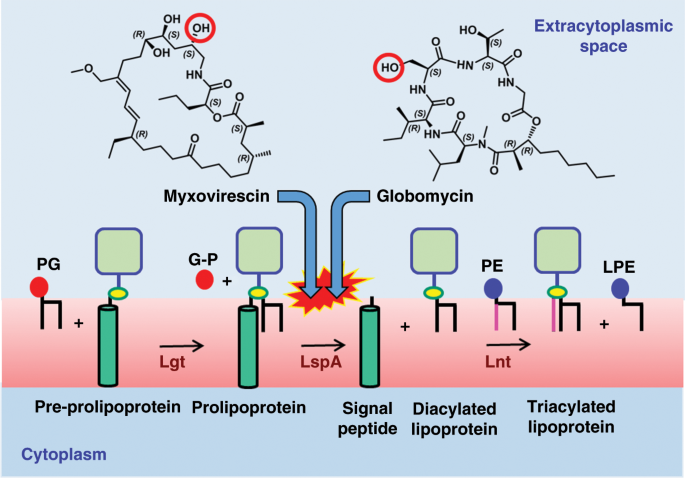
Structures of lipoprotein signal peptidase II from Staphylococcus aureus complexed with antibiotics globomycin and myxovirescin
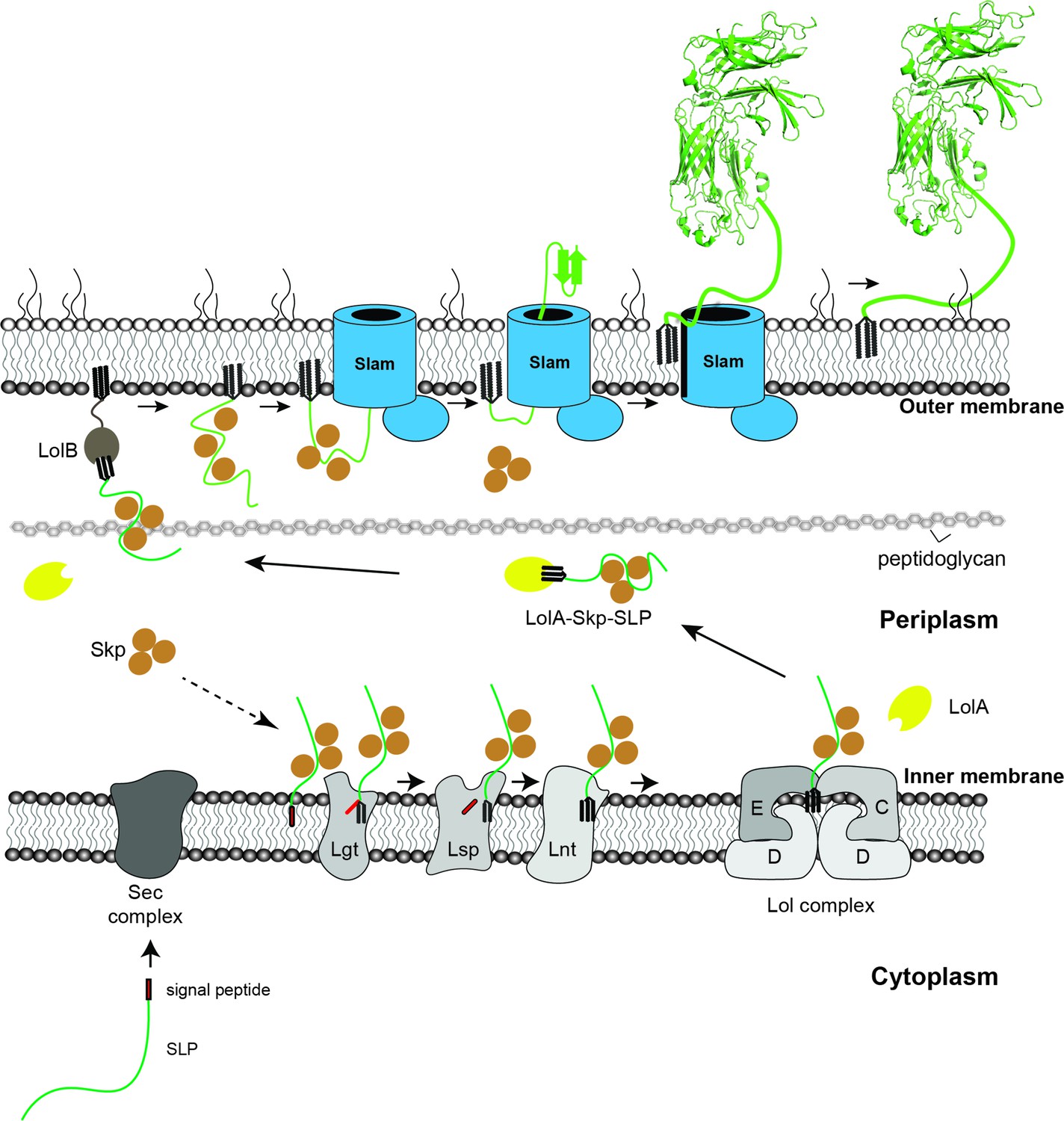
Reconstitution of surface lipoprotein translocation through the Slam translocon
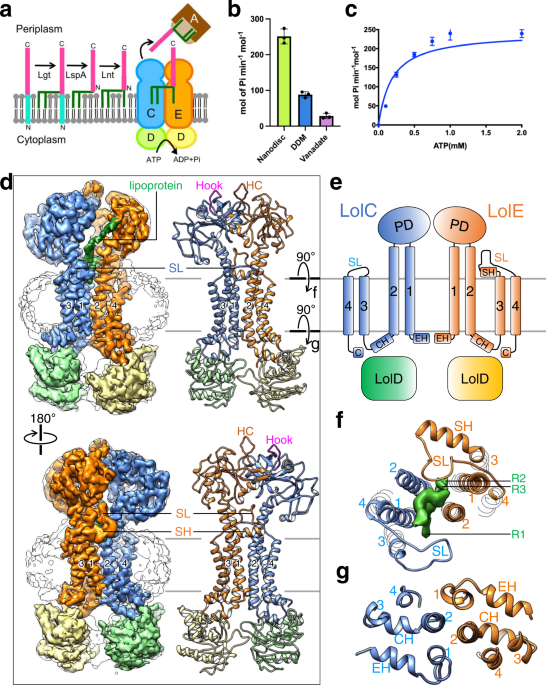
Mechanism of LolCDE as a molecular extruder of bacterial triacylated lipoproteins

Lipoproteins in Gram-negative bacteria: new insights into their biogenesis, subcellular targeting and functional roles - ScienceDirect

Mechanism of LolCDE as a molecular extruder of bacterial triacylated lipoproteins
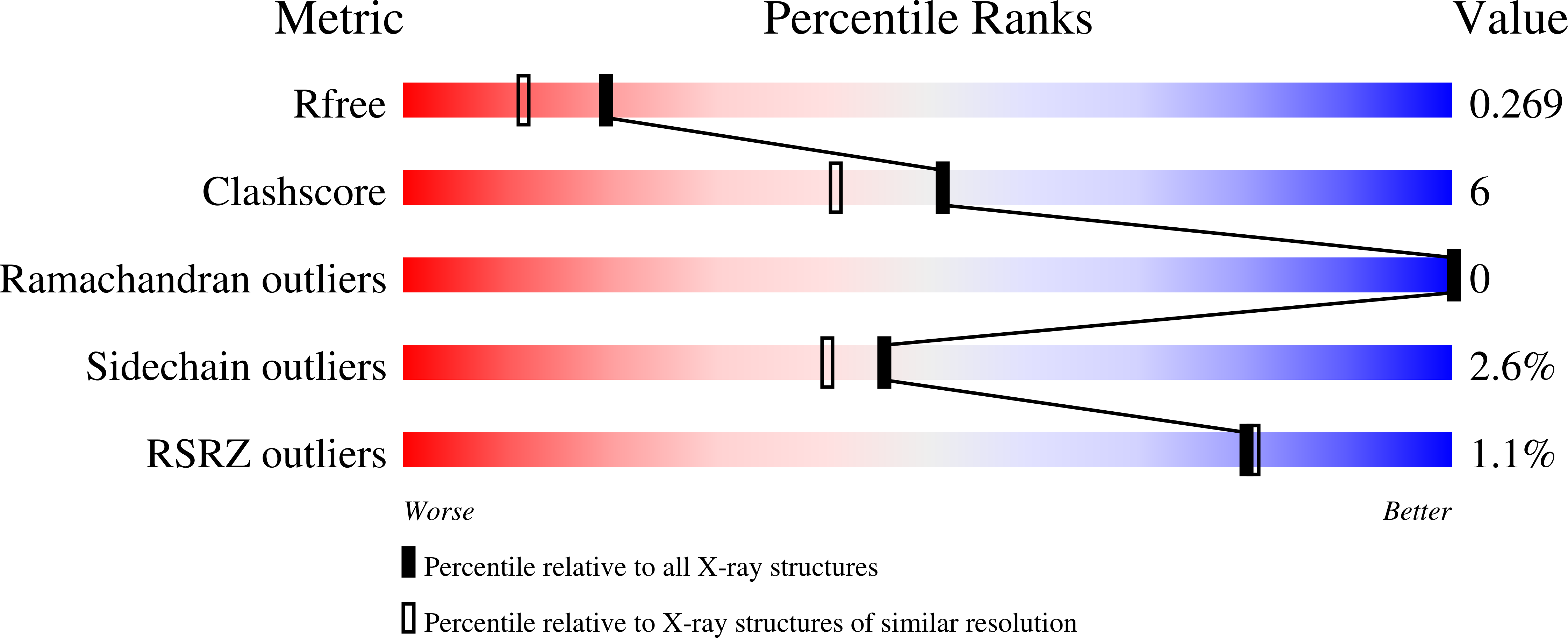
RCSB PDB - 7Z6X: Complex of E. coli LolA R43L mutant and lipoprotein
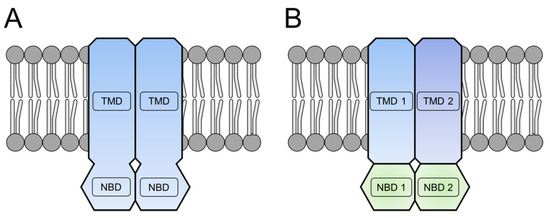
IJMS, Free Full-Text

Outer membrane lipoprotein biogenesis: Lol is not the end Philosophical Transactions of the Royal Society B: Biological Sciences

TLR2 recognition of lipoproteins and LTA. Recognition of LTA and

How to take your sticky proteins out safely
de
por adulto (o preço varia de acordo com o tamanho do grupo)







